Sigmoid vs ReLU Activation Functions: The Inference Cost of Losing a Geometric Total
A deep neural network can be understood as a geometric system, where each layer reshapes the input space to create complex decision parameters. For this to work effectively, the layers must preserve logical spatial information – specifically how far a data point is from these boundaries – as this distance allows the deeper layers to build richer, non-specific representations.
Sigmoid interrupts this process by compressing all inputs into a small range between 0 and 1. Since the values are from the decision boundaries, they are not divisible, resulting in the loss of the geometric context in all layers. This leads to poor representation and limits the depth performance.
ReLU, on the other hand, preserves the size of the positive input, allowing distance information to flow through the network. This allows deep models to continue rendering without requiring much scope or computation.
In this article, we focus on this forward behavior – we analyze how Sigmoid and ReLU differ in signal distribution and representation geometry using a two-month test, and what that means for efficiency and scalability.



Setting dependencies
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from matplotlib.colors import ListedColormap
from sklearn.datasets import make_moons
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_splitplt.rcParams.update({
"font.family": "monospace",
"axes.spines.top": False,
"axes.spines.right": False,
"figure.facecolor": "white",
"axes.facecolor": "#f7f7f7",
"axes.grid": True,
"grid.color": "#e0e0e0",
"grid.linewidth": 0.6,
})
T = {
"bg": "white",
"panel": "#f7f7f7",
"sig": "#e05c5c",
"relu": "#3a7bd5",
"c0": "#f4a261",
"c1": "#2a9d8f",
"text": "#1a1a1a",
"muted": "#666666",
}Creates a dataset
To study the effect of activation functions in a controlled setting, we first generate an artificial dataset using make_moons in scikit-learn. This creates a non-linear, two-phase problem where simple linear constraints fail, making it ideal for testing how well neural networks learn complex decisions.
We add a small amount of noise to make the task more realistic, and we set the features using StandardScaler so that both measurements are on the same scale – ensuring stable training. The dataset is then split into training and test sets to test generalization.
Finally, we visualize the data distribution. This structure serves as the basic geometry that both Sigmoid and ReLU networks will attempt to model, allowing us to later compare how each activation changes this space across layers.
X, y = make_moons(n_samples=400, noise=0.18, random_state=42)
X = StandardScaler().fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=42
)
fig, ax = plt.subplots(figsize=(7, 5))
fig.patch.set_facecolor(T["bg"])
ax.set_facecolor(T["panel"])
ax.scatter(X[y == 0, 0], X[y == 0, 1], c=T["c0"], s=40,
edgecolors="white", linewidths=0.5, label="Class 0", alpha=0.9)
ax.scatter(X[y == 1, 0], X[y == 1, 1], c=T["c1"], s=40,
edgecolors="white", linewidths=0.5, label="Class 1", alpha=0.9)
ax.set_title("make_moons -- our dataset", color=T["text"], fontsize=13)
ax.set_xlabel("x₁", color=T["muted"]); ax.set_ylabel("x₂", color=T["muted"])
ax.tick_params(colors=T["muted"]); ax.legend(fontsize=10)
plt.tight_layout()
plt.savefig("moons_dataset.png", dpi=140, bbox_inches="tight")
plt.show()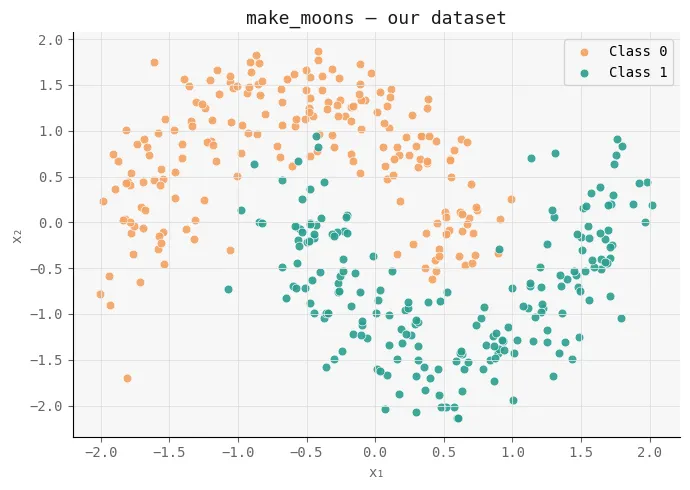

Creating a Network
Next, we use a sparse, supervised neural network to isolate the effect of activation functions. The goal here is not to build a highly optimized model, but to create a clean test setup where Sigmoid and ReLU can be compared under similar conditions.
We define both activation functions (Sigmoid and ReLU) and their derivatives, and we use the binary cross-entropy as the loss since this is a binary classification function. The TwoLayerNet class represents a simple network with 3 layers (2 hidden layers + output), where the only configurable part is the activation function.
The main detail is the initialization strategy: we use He the initialization of ReLU and Xavier the initialization of Sigmoid, we ensure that each network starts with a correct and stable state based on its activation dynamics.
The forward pass includes a layer-by-layer activation, while the backward pass performs standard gradient descent updates. Importantly, we also include diagnostic methods such as get_hidden and get_z_trace, which allow us to check how signals appear in all layers – this is important for analyzing how much geometric information is preserved or lost.
By keeping the architecture, data, and training structure constant, this implementation ensures that any differences in performance or internal representations can be directly attributed to the activation function itself – setting the stage for a clear comparison of their impact on signal propagation and resonance.
def sigmoid(z): return 1 / (1 + np.exp(-np.clip(z, -500, 500)))
def sigmoid_d(a): return a * (1 - a)
def relu(z): return np.maximum(0, z)
def relu_d(z): return (z > 0).astype(float)
def bce(y, yhat): return -np.mean(y * np.log(yhat + 1e-9) + (1 - y) * np.log(1 - yhat + 1e-9))
class TwoLayerNet:
def __init__(self, activation="relu", seed=0):
np.random.seed(seed)
self.act_name = activation
self.act = relu if activation == "relu" else sigmoid
self.dact = relu_d if activation == "relu" else sigmoid_d
# He init for ReLU, Xavier for Sigmoid
scale = lambda fan_in: np.sqrt(2 / fan_in) if activation == "relu" else np.sqrt(1 / fan_in)
self.W1 = np.random.randn(2, 8) * scale(2)
self.b1 = np.zeros((1, 8))
self.W2 = np.random.randn(8, 8) * scale(8)
self.b2 = np.zeros((1, 8))
self.W3 = np.random.randn(8, 1) * scale(8)
self.b3 = np.zeros((1, 1))
self.loss_history = []
def forward(self, X, store=False):
z1 = X @ self.W1 + self.b1; a1 = self.act(z1)
z2 = a1 @ self.W2 + self.b2; a2 = self.act(z2)
z3 = a2 @ self.W3 + self.b3; out = sigmoid(z3)
if store:
self._cache = (X, z1, a1, z2, a2, z3, out)
return out
def backward(self, lr=0.05):
X, z1, a1, z2, a2, z3, out = self._cache
n = X.shape[0]
dout = (out - self.y_cache) / n
dW3 = a2.T @ dout; db3 = dout.sum(axis=0, keepdims=True)
da2 = dout @ self.W3.T
dz2 = da2 * (self.dact(z2) if self.act_name == "relu" else self.dact(a2))
dW2 = a1.T @ dz2; db2 = dz2.sum(axis=0, keepdims=True)
da1 = dz2 @ self.W2.T
dz1 = da1 * (self.dact(z1) if self.act_name == "relu" else self.dact(a1))
dW1 = X.T @ dz1; db1 = dz1.sum(axis=0, keepdims=True)
for p, g in [(self.W3,dW3),(self.b3,db3),(self.W2,dW2),
(self.b2,db2),(self.W1,dW1),(self.b1,db1)]:
p -= lr * g
def train_step(self, X, y, lr=0.05):
self.y_cache = y.reshape(-1, 1)
out = self.forward(X, store=True)
loss = bce(self.y_cache, out)
self.backward(lr)
return loss
def get_hidden(self, X, layer=1):
"""Return post-activation values for layer 1 or 2."""
z1 = X @ self.W1 + self.b1; a1 = self.act(z1)
if layer == 1: return a1
z2 = a1 @ self.W2 + self.b2; return self.act(z2)
def get_z_trace(self, x_single):
"""Return pre-activation magnitudes per layer for ONE sample."""
z1 = x_single @ self.W1 + self.b1
a1 = self.act(z1)
z2 = a1 @ self.W2 + self.b2
a2 = self.act(z2)
z3 = a2 @ self.W3 + self.b3
return [np.abs(z1).mean(), np.abs(a1).mean(),
np.abs(z2).mean(), np.abs(a2).mean(),
np.abs(z3).mean()]Training Networks
We now train both networks under the same conditions to ensure a fair comparison. We initialize two models – one using Sigmoid and the other using ReLU – with the same random seed to start from the same weight configuration.
The training loop runs for 800 epochs using mini-batch gradient descent. At each time, we shuffle the training data, divide it into clusters, and update both networks in parallel. This setup ensures that the only dynamic change between the two operations is the activation function.
We also track losses after each period and include them periodically. This allows us to look at how each network changes over time – not just in terms of convergence speed, but whether it continues to improve or plateau.
This step is important because it establishes the first signal of difference: if both models start out the same but behave differently during training, that difference must come from the way each activation function propagates and stores information through the network.
EPOCHS = 800
LR = 0.05
BATCH = 64
net_sig = TwoLayerNet("sigmoid", seed=42)
net_relu = TwoLayerNet("relu", seed=42)
for epoch in range(EPOCHS):
idx = np.random.permutation(len(X_train))
for net in [net_sig, net_relu]:
epoch_loss = []
for i in range(0, len(idx), BATCH):
b = idx[i:i+BATCH]
loss = net.train_step(X_train[b], y_train[b], LR)
epoch_loss.append(loss)
net.loss_history.append(np.mean(epoch_loss))
if (epoch + 1) % 200 == 0:
ls = net_sig.loss_history[-1]
lr = net_relu.loss_history[-1]
print(f" Epoch {epoch+1:4d} | Sigmoid loss: {ls:.4f} | ReLU loss: {lr:.4f}")
print("n✅ Training complete.")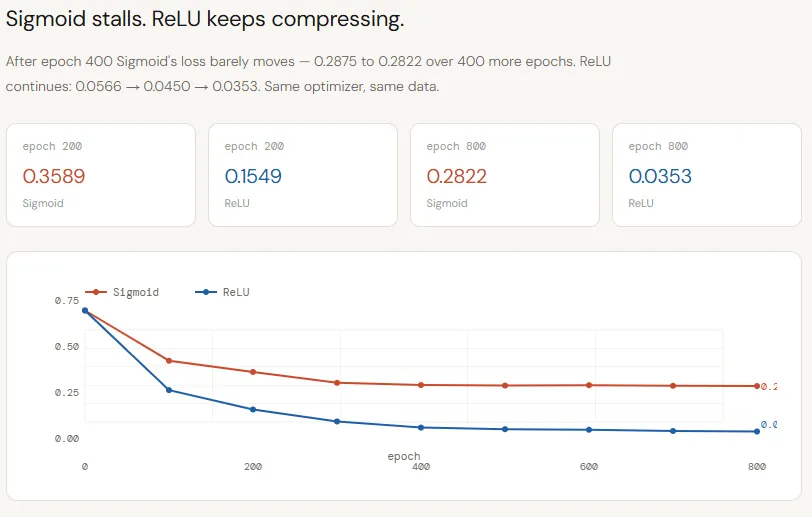

The Training Loss Curve
The loss curves make the difference between Sigmoid and ReLU very clear. Both networks start from the same initialization and are trained under the same conditions, yet their learning methods quickly diverge. The Sigmoid improves at the beginning but plateaus around ~0.28 by epoch 400, showing almost no progress after that – a sign that the network has exhausted the useful signal it can emit.
ReLU, in contrast, continues to gradually reduce the loss throughout training, dropping from ~~0.15 to ~0.03 by epoch 800. This is not just a fast convergence; shows a deeper problem: Sigmoid compression limits the flow of logical information, causing the model to stall, while ReLU preserves that signal, allowing the network to continue refining its decision boundary.
fig, ax = plt.subplots(figsize=(10, 5))
fig.patch.set_facecolor(T["bg"])
ax.set_facecolor(T["panel"])
ax.plot(net_sig.loss_history, color=T["sig"], lw=2.5, label="Sigmoid")
ax.plot(net_relu.loss_history, color=T["relu"], lw=2.5, label="ReLU")
ax.set_xlabel("Epoch", color=T["muted"])
ax.set_ylabel("Binary Cross-Entropy Loss", color=T["muted"])
ax.set_title("Training Loss -- same architecture, same init, same LRnonly the activation differs",
color=T["text"], fontsize=12)
ax.legend(fontsize=11)
ax.tick_params(colors=T["muted"])
# Annotate final losses
for net, color, va in [(net_sig, T["sig"], "bottom"), (net_relu, T["relu"], "top")]:
final = net.loss_history[-1]
ax.annotate(f" final: {final:.4f}", xy=(EPOCHS-1, final),
color=color, fontsize=9, va=va)
plt.tight_layout()
plt.savefig("loss_curves.png", dpi=140, bbox_inches="tight")
plt.show()

Decision Boundary Episodes
Visualization of the decision boundary makes the difference even more apparent. The Sigmoid network learns about the linear boundary, failing to capture the curved structure of the two-month dataset, resulting in low accuracy (~79%). This is a direct consequence of its compressed internal representation — the network does not have enough geometric signal to construct a complex boundary.
In contrast, the ReLU network learns a highly nonlinear, well-adapted boundary that closely follows the data distribution, achieving a very high accuracy (~96%). Because ReLU preserves dimensions across layers, it enables the network to continuously bend and refine the decision space, turning depth into real expressive power instead of wasted capacity.
def plot_boundary(ax, net, X, y, title, color):
h = 0.025
x_min, x_max = X[:, 0].min() - 0.5, X[:, 0].max() + 0.5
y_min, y_max = X[:, 1].min() - 0.5, X[:, 1].max() + 0.5
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
grid = np.c_[xx.ravel(), yy.ravel()]
Z = net.forward(grid).reshape(xx.shape)
# Soft shading
cmap_bg = ListedColormap(["#fde8c8", "#c8ece9"])
ax.contourf(xx, yy, Z, levels=50, cmap=cmap_bg, alpha=0.85)
ax.contour(xx, yy, Z, levels=[0.5], colors=[color], linewidths=2)
ax.scatter(X[y==0, 0], X[y==0, 1], c=T["c0"], s=35,
edgecolors="white", linewidths=0.4, alpha=0.9)
ax.scatter(X[y==1, 0], X[y==1, 1], c=T["c1"], s=35,
edgecolors="white", linewidths=0.4, alpha=0.9)
acc = ((net.forward(X) >= 0.5).ravel() == y).mean()
ax.set_title(f"{title}nTest acc: {acc:.1%}", color=color, fontsize=12)
ax.set_xlabel("x₁", color=T["muted"]); ax.set_ylabel("x₂", color=T["muted"])
ax.tick_params(colors=T["muted"])
fig, axes = plt.subplots(1, 2, figsize=(13, 5.5))
fig.patch.set_facecolor(T["bg"])
fig.suptitle("Decision Boundaries learned on make_moons",
fontsize=13, color=T["text"])
plot_boundary(axes[0], net_sig, X_test, y_test, "Sigmoid", T["sig"])
plot_boundary(axes[1], net_relu, X_test, y_test, "ReLU", T["relu"])
plt.tight_layout()
plt.savefig("decision_boundaries.png", dpi=140, bbox_inches="tight")
plt.show()

Layer-by-Layer Signal Trace
This chart tracks how the signal changes in all layers from a point away from the decision boundary – and clearly shows where the Sigmoid fails. Both networks start with the same pre-activation size in the first layer (~2.0), but Sigmoid quickly compresses to ~0.3, while ReLU maintains a higher value. As we go deeper, the Sigmoid continues to spread the signal into a smaller band (0.5–0.6), effectively erasing the logical difference. ReLU, on the other hand, maintains and increases the size, with the last layer reaching values up to 9-20.
This means that the output neuron in the ReLU network makes decisions based on a strong, well-differentiated signal, while the Sigmoid network is forced to differentiate using a weak, suppressed one. The key takeaway is that ReLU maintains a distance from the decision boundary in all layers, allowing that information to be integrated, while Sigmoid gradually destroys it.
far_class0 = X_train[y_train == 0][np.argmax(
np.linalg.norm(X_train[y_train == 0] - [-1.2, -0.3], axis=1)
)]
far_class1 = X_train[y_train == 1][np.argmax(
np.linalg.norm(X_train[y_train == 1] - [1.2, 0.3], axis=1)
)]
stage_labels = ["z₁ (pre)", "a₁ (post)", "z₂ (pre)", "a₂ (post)", "z₃ (out)"]
x_pos = np.arange(len(stage_labels))
fig, axes = plt.subplots(1, 2, figsize=(13, 5.5))
fig.patch.set_facecolor(T["bg"])
fig.suptitle("Layer-by-layer signal magnitude -- a point far from the boundary",
fontsize=12, color=T["text"])
for ax, sample, title in zip(
axes,
[far_class0, far_class1],
["Class 0 sample (deep in its moon)", "Class 1 sample (deep in its moon)"]
):
ax.set_facecolor(T["panel"])
sig_trace = net_sig.get_z_trace(sample.reshape(1, -1))
relu_trace = net_relu.get_z_trace(sample.reshape(1, -1))
ax.plot(x_pos, sig_trace, "o-", color=T["sig"], lw=2.5, markersize=8, label="Sigmoid")
ax.plot(x_pos, relu_trace, "s-", color=T["relu"], lw=2.5, markersize=8, label="ReLU")
for i, (s, r) in enumerate(zip(sig_trace, relu_trace)):
ax.text(i, s - 0.06, f"{s:.3f}", ha="center", fontsize=8, color=T["sig"])
ax.text(i, r + 0.04, f"{r:.3f}", ha="center", fontsize=8, color=T["relu"])
ax.set_xticks(x_pos); ax.set_xticklabels(stage_labels, color=T["muted"], fontsize=9)
ax.set_ylabel("Mean |activation|", color=T["muted"])
ax.set_title(title, color=T["text"], fontsize=11)
ax.tick_params(colors=T["muted"]); ax.legend(fontsize=10)
plt.tight_layout()
plt.savefig("signal_trace.png", dpi=140, bbox_inches="tight")
plt.show()

Hidden Space Scatter
This is a very important visualization because it reveals exactly how each network uses (or fails to use) depth. In the Sigmoid network (left), both classes fall into a strong, overlapping area – a diagonal smear where points are closely related. The standard deviation actually decreases from layer 1 (0.26) to layer 2 (0.19), which means that the representation decreases with depth. Each layer compresses the signal further, stripping the spatial structure needed to separate the classes.
ReLU shows the opposite behavior. In layer 1, while some neurons are inactive (“dead zone”), the active ones are already spread over a wide range (1.15 std), indicating conserved variation. In layer 2, this extends even further (1.67 std), and the classes are clearly distinguishable – one is pushed to the upper range of activation while the other stays close to zero. At this point, the function of the output layer is irrelevant.
fig, axes = plt.subplots(2, 2, figsize=(13, 10))
fig.patch.set_facecolor(T["bg"])
fig.suptitle("Hidden-space representations on make_moons test set",
fontsize=13, color=T["text"])
for col, (net, color, name) in enumerate([
(net_sig, T["sig"], "Sigmoid"),
(net_relu, T["relu"], "ReLU"),
]):
for row, layer in enumerate([1, 2]):
ax = axes[row][col]
ax.set_facecolor(T["panel"])
H = net.get_hidden(X_test, layer=layer)
ax.scatter(H[y_test==0, 0], H[y_test==0, 1], c=T["c0"], s=40,
edgecolors="white", linewidths=0.4, alpha=0.85, label="Class 0")
ax.scatter(H[y_test==1, 0], H[y_test==1, 1], c=T["c1"], s=40,
edgecolors="white", linewidths=0.4, alpha=0.85, label="Class 1")
spread = H.std()
ax.text(0.04, 0.96, f"std: {spread:.4f}",
transform=ax.transAxes, fontsize=9, va="top",
color=T["text"],
bbox=dict(boxstyle="round,pad=0.3", fc="white", ec=color, alpha=0.85))
ax.set_title(f"{name} -- Layer {layer} hidden space",
color=color, fontsize=11)
ax.set_xlabel(f"Unit 1", color=T["muted"])
ax.set_ylabel(f"Unit 2", color=T["muted"])
ax.tick_params(colors=T["muted"])
if row == 0 and col == 0: ax.legend(fontsize=9)
plt.tight_layout()
plt.savefig("hidden_space.png", dpi=140, bbox_inches="tight")
plt.show()



Check it out Full Codes here. Also, feel free to follow us Twitter and don’t forget to join our 120k+ ML SubReddit and Subscribe to Our newspaper. Wait! are you on telegram? now you can join us on telegram too.
Need to work with us on developing your GitHub Repo OR Hug Face Page OR Product Release OR Webinar etc.? contact us

I am a Civil Engineering Graduate (2022) from Jamia Millia Islamia, New Delhi, and I am very interested in Data Science, especially Neural Networks and its application in various fields.



